Representative sets exclude sequence homologs sharing > 25% amino acid identity. SALIGN įold classification based on structure-structure alignment of proteins (FSSP) FSSP is based on a comprehensive comparison of PDB proteins (greater than 30 amino acids in length) using DALI. The algorithm uses the three-dimensional coordinates of each protein to calculate distance matrices comparing residues. Low values of RMSD mean similar structuresĭali (Distance mAtrix aLIgnment) DALI offers pairwise alignments of protein structures. The RMSD of two aligned structures indicates their divergence from one another. The outputs of a structural alignment are a superposition of the atomic coordinates and a minimal Root Mean Square Distance (RMSD) between the structures. Structure Alignments There are many different algorithms for structural Alignment. Sequence and Structure alignments of two Retinol Binding Protein 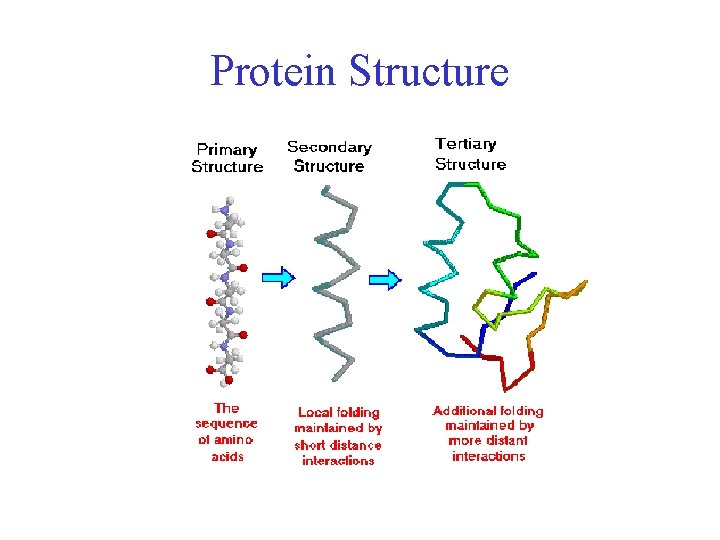
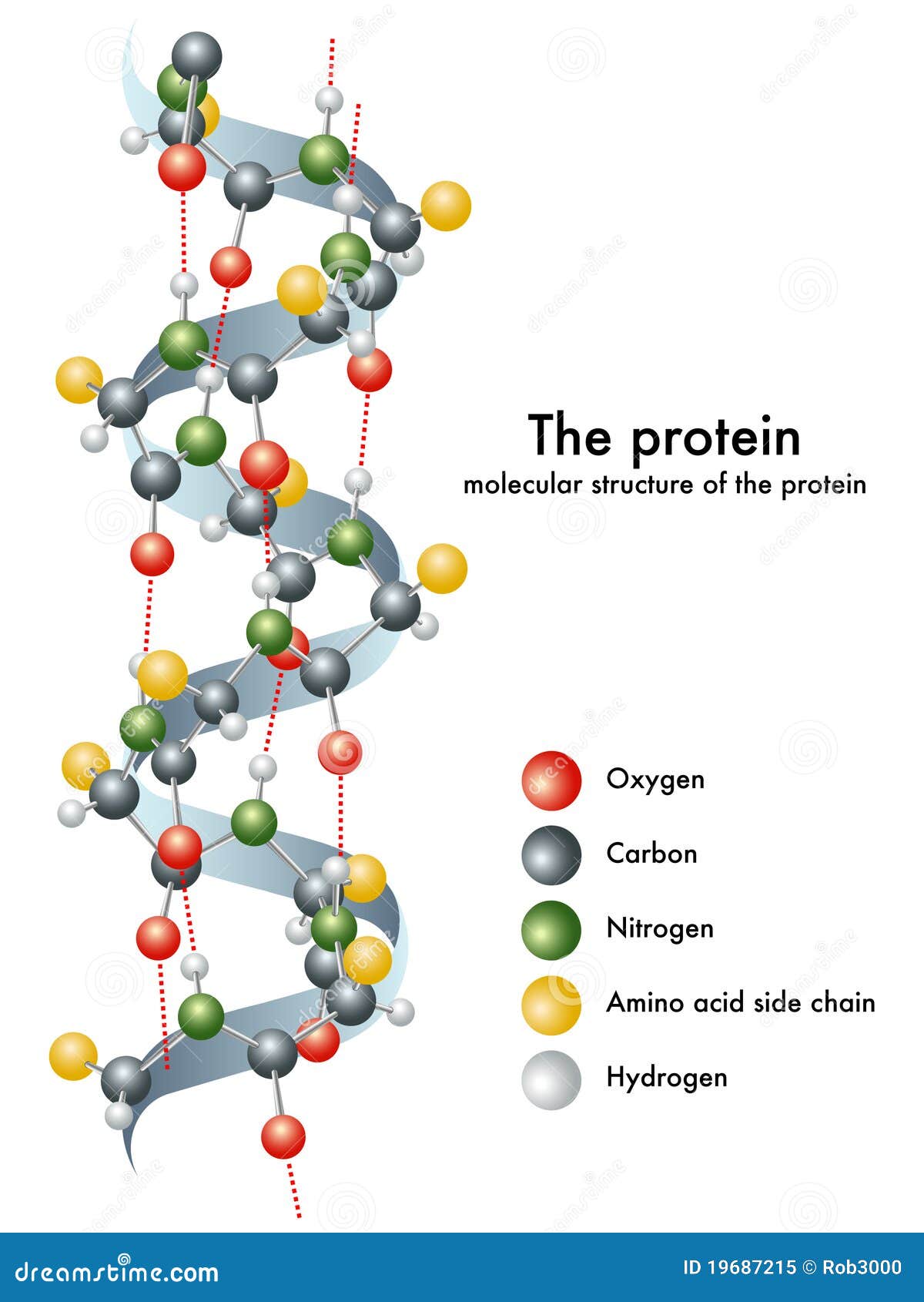
Comparative modeling (homology) Based on structural homology īased on Sequence homology Comparative Modeling Similar sequences suggests similar structure.Predicting 3D Structure Outstanding difficult problem Based on sequence homology Open file “1aqb” (PDB coordinate file).Launch a 3D viewer program For example we will use the program Pymol The program can be downloaded freely from the Pymol homepage.Download the coordinates of the structure from the PDB.Tertiary: the 3D shape of the fully folded polypeptide chain Secondary: -helices, β-sheets and loops. The Different levels of Protein Structure Primary: amino acid linear sequence. Protein Tertiary Structure Prediction Structural Bioinformatics 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
